At the end of September, the Journal of Clinical Microbiology published a remarkable study evaluating the accuracy and robustness of fully automated Rapid Antimicrobial Susceptibility Testing (RAST) directly from positive blood cultures (BCs). This paper – by A. Chekraoui and colleagues – will significantly impact the whole microbiology community, as the spread of multidrug-resistant organisms and the increase of resistant infections make RAST more valuable than ever. Let’s see what it’s all about!
Although many positive BCs’ RAST methods have been released and rapid standardized disk-diffusion methods have been developed by EUCAST to enable susceptibility reports within 4 to 8 h and 16 to 18 h from positive BCs, the labor-intensive manual setup and the strictly defined reading time points have hampered their large-scale use in clinical microbiology laboratories. Nevertheless, shorter times for delivering accurate antimicrobial susceptibility results to provide patients with a rapid antimicrobial treatment remain one of the primary objectives to be achieved by microbiologists.
Here is where lab automation comes to help. In recent years, full laboratory automation has permitted the flexible, cost-effective, and accurate disk-diffusion method to return to the center stage and strengthen its position among other AST methods currently used in clinical microbiology laboratories.
This study evaluates (and, spoiler alert, confirms) the accuracy and robustness of the fully automated RAST in providing fast and accurate results.
Briefly, the study was conducted in two phases.
In the first one, researchers spiked negative BCs bottles with 779 non-duplicate bacterial isolates, including various resistant phenotypes. They paid particular attention to testing a large number of clinical isolates expressing diverse resistant phenotypes, including Escherichia coli, Klebsiella pneumoniae, Pseudomonas aeruginosa, Acinetobacter baumannii, Staphylococcus aureus, Enterococcus faecalis, and Enterococcus faecium.
In the second phase, the researchers performed a prospective clinical trial on more than 500 positive BCs processed in routine at the Bacteriology Laboratory of Geneva University Hospitals.
In both cases, fully automated disk-diffusion RAST plate preparation and S/I/R inhibition zone interpretation were performed by WASPLab and Radian at 4, 6, and 8 hrs. The results, validated by expert technologists, were assessed against EUCAST standardized disk-diffusion testing results.
First, Radian interpretation demonstrated great accuracy, as the manual adjustment of the inhibition zone diameter concerned only about 5% of all tested disks.
Second, the results of the spiked BCs precisely predicted the results of the clinical trial: all antibiotics tested could be released at 4 h with high confidence, with a categorical agreement for RAST compared with the EUCAST standardized disk-diffusion test results greater than 95% for all the antibiotics tested – except for amikacin, released at 6 hrs. The only organism that posed some challenges was P. aeruginosa, and greater attention should be paid to interpreting this organism with the RAST technique.
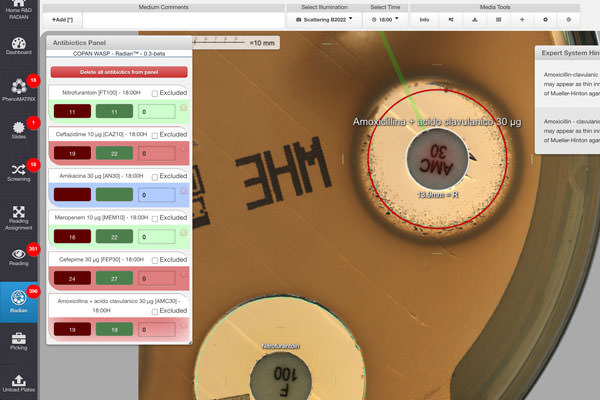
In a previous paper, Dr. Cherkaoui’s team validated Radian’s full automation of standardized disk-diffusion AST. Now, they extended the use of Radian’s fully automated solution to the EUCAST RAST method directly from positive BCs. The group established that the performance of fully automated RAST from positive BCs is robust even for detecting ESBL, carbapenemase-producing bacteria, and MRSA. Moreover, the automation enhanced the percentage of readable inhibition zones and reduced the rate of isolates categorized with technical uncertainty.
In other words, this study demonstrates that fully automated EUCAST RAST can substantially improve laboratory workflow by reducing hands-on time and removing the strong constraints linked to manual read-outs at precisely defined times, empowering clinicians to provide prompt treatment indications to patients.
If you want to explore the specific results for each organism and antibiotic, we suggest you read the full paper below: it’s Open access and contains clear tables to be consulted.

Write us to satisfy your curiosity, get information or start a great collaboration!
Use our search tool and be surprised by how easy it is to get right to the point.